skrub.GapEncoder#
Usage examples at the bottom of this page.
- class skrub.GapEncoder(n_components=10, batch_size=1024, gamma_shape_prior=1.1, gamma_scale_prior=1.0, rho=0.95, rescale_rho=False, hashing=False, hashing_n_features=4096, init='k-means++', max_iter=5, ngram_range=(2, 4), analyzer='char', add_words=False, random_state=None, rescale_W=True, max_iter_e_step=1, max_no_improvement=5, verbose=0)[source]#
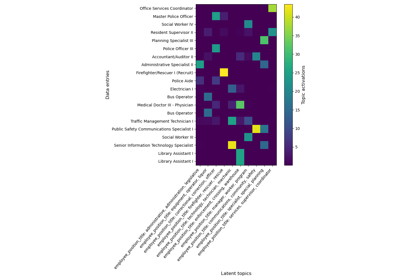
Constructs latent topics with continuous encoding.
Note
GapEncoderis a type of single-column transformer. Unlike most scikit-learn estimators, itsfit,transformandfit_transformmethods expect a single column (a pandas or polars Series) rather than a full dataframe. To apply this transformer to one or more columns in a dataframe, use it as a parameter in askrub.TableVectorizerorsklearn.compose.ColumnTransformer. In theColumnTransformer, pass a single column:make_column_transformer((GapEncoder(), 'col_name_1'), (GapEncoder(), 'col_name_2'))instead ofmake_column_transformer((GapEncoder(), ['col_name_1', 'col_name_2'])).This encoder can be understood as a continuous encoding on a set of latent categories estimated from the data. The latent categories are built by capturing combinations of substrings that frequently co-occur.
The GapEncoder supports online learning on batches of data for scalability through the GapEncoder.partial_fit method.
The principle is as follows:
Given an input string array X, we build its bag-of-n-grams representation V (n_samples, vocab_size).
Instead of using the n-grams counts as encodings, we look for low- dimensional representations by modeling n-grams counts as linear combinations of topics
V = HW, with W (n_topics, vocab_size) the topics and H (n_samples, n_topics) the associated activations.Assuming that n-grams counts follow a Poisson law, we fit H and W to maximize the likelihood of the data, with a Gamma prior for the activations H to induce sparsity.
In practice, this is equivalent to a non-negative matrix factorization with the Kullback-Leibler divergence as loss, and a Gamma prior on H. We thus optimize H and W with the multiplicative update method.
“Gap” stands for “Gamma-Poisson”, the families of distributions that are used to model the importance of topics in a document (Gamma), and the term frequencies in a document (Poisson).
Input columns that do not have a string or Categorical dtype are rejected by raising a
RejectColumnexception.- Parameters:
- n_components
int, optional, default=10 Number of latent categories used to model string data.
- batch_size
int, optional, default=1024 Number of samples per batch.
- gamma_shape_prior
float, optional, default=1.1 Shape parameter for the Gamma prior distribution.
- gamma_scale_prior
float, optional, default=1.0 Scale parameter for the Gamma prior distribution.
- rho
float, optional, default=0.95 Weight parameter for the update of the W matrix.
- rescale_rho
bool, optional, default=False If True, use
rho ** (batch_size / len(X))instead of rho to obtain an update rate per iteration that is independent of the batch size.- hashing
bool, optional, default=False If True, HashingVectorizer is used instead of CountVectorizer. It has the advantage of being very low memory, scalable to large datasets as there is no need to store a vocabulary dictionary in memory.
- hashing_n_features
int, default=2**12 Number of features for the HashingVectorizer. Only relevant if hashing=True.
- init{‘k-means++’, ‘random’, ‘k-means’}, default=’k-means++’
Initialization method of the W matrix. If init=’k-means++’, we use the init method of KMeans. If init=’random’, topics are initialized with a Gamma distribution. If init=’k-means’, topics are initialized with a KMeans on the n-grams counts.
- max_iter
int, default=5 Maximum number of iterations on the input data.
- ngram_range
int2-tuple, default=(2, 4) - The lower and upper boundaries of the range of n-values for different
n-grams used in the string similarity. All values of n such that
min_n <= n <= max_nwill be used.
- analyzer{‘word’, ‘char’, ‘char_wb’}, default=’char’
Analyzer parameter for the HashingVectorizer / CountVectorizer. Describes whether the matrix V to factorize should be made of word counts or character-level n-gram counts. Option ‘char_wb’ creates character n-grams only from text inside word boundaries; n-grams at the edges of words are padded with space.
- add_words
bool, default=False If True, add the words counts to the bag-of-n-grams representation of the input data.
- random_state
intorRandomState, optional Random number generator seed for reproducible output across multiple function calls.
- rescale_W
bool, default=True If True, the weight matrix W is rescaled at each iteration to have a l1 norm equal to 1 for each row.
- max_iter_e_step
int, default=1 Maximum number of iterations to adjust the activations h at each step.
- max_no_improvement
int, default=5 Control early stopping based on the consecutive number of mini batches that do not yield an improvement on the smoothed cost function. To disable early stopping and run the process fully, set
max_no_improvement=None.- verbose
int, default=0 Verbosity level. The higher, the more granular the logging.
- n_components
See also
MinHashEncoderEncode string columns as a numeric array with the minhash method.
SimilarityEncoderEncode string columns as a numeric array with n-gram string similarity.
deduplicateDeduplicate data by hierarchically clustering similar strings.
References
For a detailed description of the method, see Encoding high-cardinality string categorical variables by Cerda, Varoquaux (2019).
Examples
>>> import pandas as pd >>> from skrub import GapEncoder >>> enc = GapEncoder(n_components=2, random_state=0)
Let’s encode the following non-normalized data:
>>> X = pd.Series(['Paris, FR', 'Paris', 'London, UK', 'Paris, France', ... 'london', 'London, England', 'London', 'Pqris'], name='city') >>> enc.fit(X) GapEncoder(n_components=2, random_state=0)
The GapEncoder has found the following two topics:
>>> enc.get_feature_names_out() ['city: england, london, uk', 'city: france, paris, pqris']
It got it right, reccuring topics are “London” and “England” on the one side and “Paris” and “France” on the other.
As this is a continuous encoding, we can look at the level of activation of each topic for each category:
>>> enc.transform(X) city: england, london, uk city: france, paris, pqris 0 0.051816 10.548184 1 0.050134 4.549866 2 12.046517 0.053483 3 0.052270 16.547730 4 6.049970 0.050030 5 19.545227 0.054773 6 6.049970 0.050030 7 0.060120 4.539880
The higher the value, the bigger the correspondence with the topic.
- Attributes:
Methods
fit(X[, y])Fit the GapEncoder on X.
fit_transform(X[, y])Fit to data, then transform it.
get_feature_names_out([n_labels])Return the labels that best summarize the learned components/topics.
Get metadata routing of this object.
get_params([deep])Get parameters for this estimator.
partial_fit(X[, y])Partial fit this instance on X.
score(X)Score this instance of X.
set_fit_request(*[, column])Request metadata passed to the
fitmethod.set_output(*[, transform])Set output container.
set_params(**params)Set the parameters of this estimator.
transform(X)Return the encoded vectors (activations) H of input strings in X.
- fit(X, y=None)[source]#
Fit the GapEncoder on X.
- Parameters:
- XColumn, shape (n_samples, )
The string data to fit the model on.
- y
None Unused, only here for compatibility.
- Returns:
- GapEncoderColumn
The fitted GapEncoderColumn instance (self).
- fit_transform(X, y=None, **fit_params)[source]#
Fit to data, then transform it.
Fits transformer to X and y with optional parameters fit_params and returns a transformed version of X.
- Parameters:
- Xarray_like of shape (n_samples, n_features)
Input samples.
- yarray_like of shape (n_samples,) or (n_samples, n_outputs), default=None
Target values (None for unsupervised transformations).
- **fit_params
dict Additional fit parameters.
- Returns:
- get_feature_names_out(n_labels=3)[source]#
Return the labels that best summarize the learned components/topics.
For each topic, labels with the highest activations are selected.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- partial_fit(X, y=None)[source]#
Partial fit this instance on X.
To be used in an online learning procedure where batches of data are coming one by one.
- Parameters:
- XColumn, shape (n_samples, )
The string data to fit the model on.
- y
None Unused, only here for compatibility.
- Returns:
- GapEncoderColumn
The fitted GapEncoderColumn instance (self).
- score(X)[source]#
Score this instance of X.
Returns the Kullback-Leibler divergence between the n-grams counts matrix V of X, and its non-negative factorization HW.
- Parameters:
- XColumn, shape (n_samples, )
The data to encode.
- Returns:
floatThe Kullback-Leibler divergence.
- set_fit_request(*, column='$UNCHANGED$')[source]#
Request metadata passed to the
fitmethod.Note that this method is only relevant if
enable_metadata_routing=True(seesklearn.set_config()). Please see User Guide on how the routing mechanism works.The options for each parameter are:
True: metadata is requested, and passed tofitif provided. The request is ignored if metadata is not provided.False: metadata is not requested and the meta-estimator will not pass it tofit.None: metadata is not requested, and the meta-estimator will raise an error if the user provides it.str: metadata should be passed to the meta-estimator with this given alias instead of the original name.
The default (
sklearn.utils.metadata_routing.UNCHANGED) retains the existing request. This allows you to change the request for some parameters and not others.Added in version 1.3.
Note
This method is only relevant if this estimator is used as a sub-estimator of a meta-estimator, e.g. used inside a
Pipeline. Otherwise it has no effect.
- set_output(*, transform=None)[source]#
Set output container.
See Introducing the set_output API for an example on how to use the API.
- Parameters:
- transform{“default”, “pandas”, “polars”}, default=None
Configure output of transform and fit_transform.
“default”: Default output format of a transformer
“pandas”: DataFrame output
“polars”: Polars output
None: Transform configuration is unchanged
Added in version 1.4: “polars” option was added.
- Returns:
- selfestimator instance
Estimator instance.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **params
dict Estimator parameters.
- **params
- Returns:
- selfestimator instance
Estimator instance.
Examples using skrub.GapEncoder#
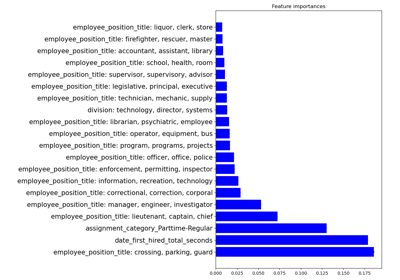
Encoding: from a dataframe to a numerical matrix for machine learning